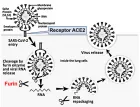
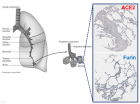
Figure 3
Reasons why new coronavirus, SARS-CoV-2 infections are likely to spread
Takuma Hayashi*, Takashi Ura, Kaoru Abiko, Masaki Mandan, Nobuo Yaegashi and Ikuo Konishi
Published: 28 April, 2020 | Volume 3 - Issue 1 | Pages: 001-003
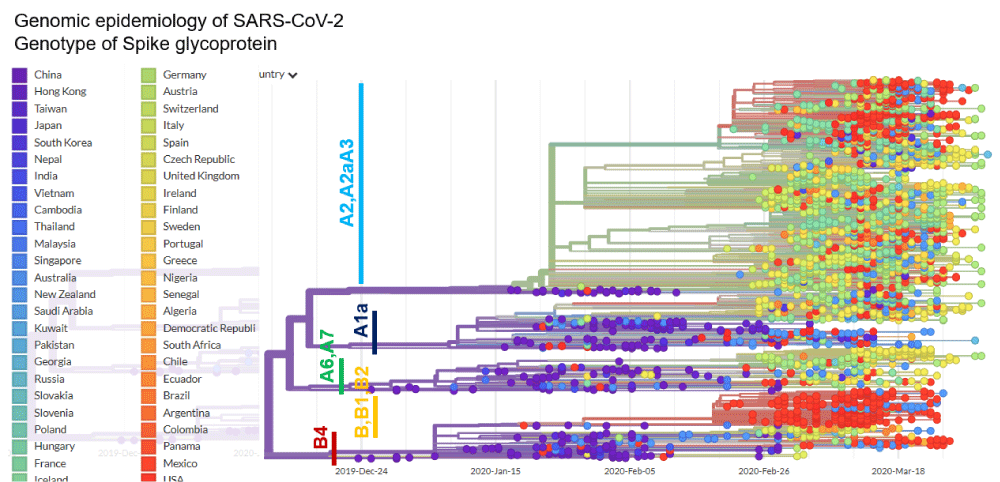
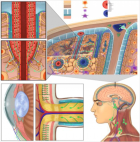
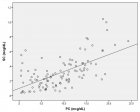
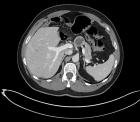
Figure 3:
Phylogenetic network analysis of amino acid sequences of spike glycoprotein derived from 160 complete human SARS-Cov-2 genomes. Our research results find ten central variants distinguished by amino acid changes, which we have named A1a, A2, A2a, A3, A6, A7, B, B1, B2, and B4, with being the ancestral type according to the bat out group coronavirus. The A1a, A6, A7 and B4 types are found in significant proportions in East Asia, A2, A2a, and A3 types are found in Europeans, B, B1, and B2 types are found in Americans. In contrast, the B4 type is the most common type in America and East Asia.
Read Full Article HTML DOI: 10.29328/journal.jgmgt.1001005 Cite this Article Read Full Article PDF
More Images
Similar Articles
-
Reasons why new coronavirus, SARS-CoV-2 infections are likely to spreadTakuma Hayashi*,Takashi Ura,Kaoru Abiko,Masaki Mandan,Nobuo Yaegashi,Ikuo Konishi. Reasons why new coronavirus, SARS-CoV-2 infections are likely to spread. . 2020 doi: 10.29328/journal.jgmgt.1001005; 3: 001-003
Recently Viewed
-
The Urinary Microbiome: Shifting Paradigms from Sterile Urine to Microbial Dysbiosis in Chronic Pelvic Pain SyndromeMohammed Amine Elafari*,Mohammed Amine Bibat,Mamad Ayoub,Amine Slaoui,Tarik Karmouni,Abdelatif Koutani,Khalid Elkhader. The Urinary Microbiome: Shifting Paradigms from Sterile Urine to Microbial Dysbiosis in Chronic Pelvic Pain Syndrome. Int J Clin Microbiol Biochem Technol. 2026: doi: 10.29328/journal.ijcmbt.1001036; 9: 022-026
-
Unusual Complications of a Dental Prosthesis Esophageal Foreign Body: About a CaseRichard Edward Alain Deguenonvo,Ndèye Fatou Thiam*,Mouhamadou Diouldé Diallo,Abdou Sy,Amadou Thiam,Abdoulaye Diop,Mame Sanou Diouf,Baye Karim Diallo. Unusual Complications of a Dental Prosthesis Esophageal Foreign Body: About a Case. Adv Treat ENT Disord. 2025: doi: 10.29328/journal.ated.1001016; 9: 001-004
-
One-time CRISPR Adenine Base Editing Intervention in SMA: From SMN2 Splice Correction to Motor Neuron RescueSheena P Kochumon,Najma Nujoom,Prem Jagadeesan,Vinod Scaria,DM Vasudevan,KP Soman,Cherupally Krishnan Krishnan Nair*. One-time CRISPR Adenine Base Editing Intervention in SMA: From SMN2 Splice Correction to Motor Neuron Rescue. J Genet Med Gene Ther. 2026: doi: 10.29328/journal.jgmgt.1001014; 9: 1-6
-
Germline BRCA1 Mutation inSquamous Cell Carcinoma of Oesophagus: Driver versus Passenger MutationAmrit Kaur Kaler*, Shraddha Manoj Upadhyay, Nandini Shyamali Bora, Ankita Nikam, Kavya P, Nivetha Athikeri, Dattatray B Solanki, Imran Shaikh, Rajesh Mistry. Germline BRCA1 Mutation inSquamous Cell Carcinoma of Oesophagus: Driver versus Passenger Mutation. J Genet Med Gene Ther. 2024: doi: 10.29328/journal.jgmgt.1001011; 7: 015-019
-
The Role of Genetic Mutations in the HPGD & SLCO2A1 Genes in Pachydermoperiostosis SyndromeShahin Asadi*,Arezo Zare,Sima Koohestani. The Role of Genetic Mutations in the HPGD & SLCO2A1 Genes in Pachydermoperiostosis Syndrome. J Genet Med Gene Ther. 2025: doi: 10.29328/journal.jgmgt.1001013; 8: 001-005
Most Viewed
-
Physical Performance in the Overweight/Obesity Children Evaluation and RehabilitationCristina Popescu, Mircea-Sebastian Șerbănescu, Gigi Calin*, Magdalena Rodica Trăistaru. Physical Performance in the Overweight/Obesity Children Evaluation and Rehabilitation. Ann Clin Endocrinol Metabol. 2024 doi: 10.29328/journal.acem.1001030; 8: 004-012
-
Hypercalcaemic Crisis Associated with Hyperthyroidism: A Rare and Challenging PresentationKarthik Baburaj*, Priya Thottiyil Nair, Abeed Hussain, Vimal MV. Hypercalcaemic Crisis Associated with Hyperthyroidism: A Rare and Challenging Presentation. Ann Clin Endocrinol Metabol. 2024 doi: 10.29328/journal.acem.1001029; 8: 001-003
-
Effects of dietary supplementation on progression to type 2 diabetes in subjects with prediabetes: a single center randomized double-blind placebo-controlled trialSathit Niramitmahapanya*,Preeyapat Chattieng,Tiersidh Nasomphan,Korbtham Sathirakul. Effects of dietary supplementation on progression to type 2 diabetes in subjects with prediabetes: a single center randomized double-blind placebo-controlled trial. Ann Clin Endocrinol Metabol. 2023 doi: 10.29328/journal.acem.1001026; 7: 00-007
-
Exceptional cancer responders: A zone-to-goDaniel Gandia,Cecilia Suárez*. Exceptional cancer responders: A zone-to-go. Arch Cancer Sci Ther. 2023 doi: 10.29328/journal.acst.1001033; 7: 001-002
-
Ectopic adrenal tissue at the spermatic cordAbdallah Chaachou,Nizar Cherni,Wael Ferjaoui*,Mohamed Dridi,Samir Ghozzi. Ectopic adrenal tissue at the spermatic cord. J Clin Med Exp Images. 2022 doi: 10.29328/journal.jcmei.1001024; 6: 001-002

If you are already a member of our network and need to keep track of any developments regarding a question you have already submitted, click "take me to my Query."